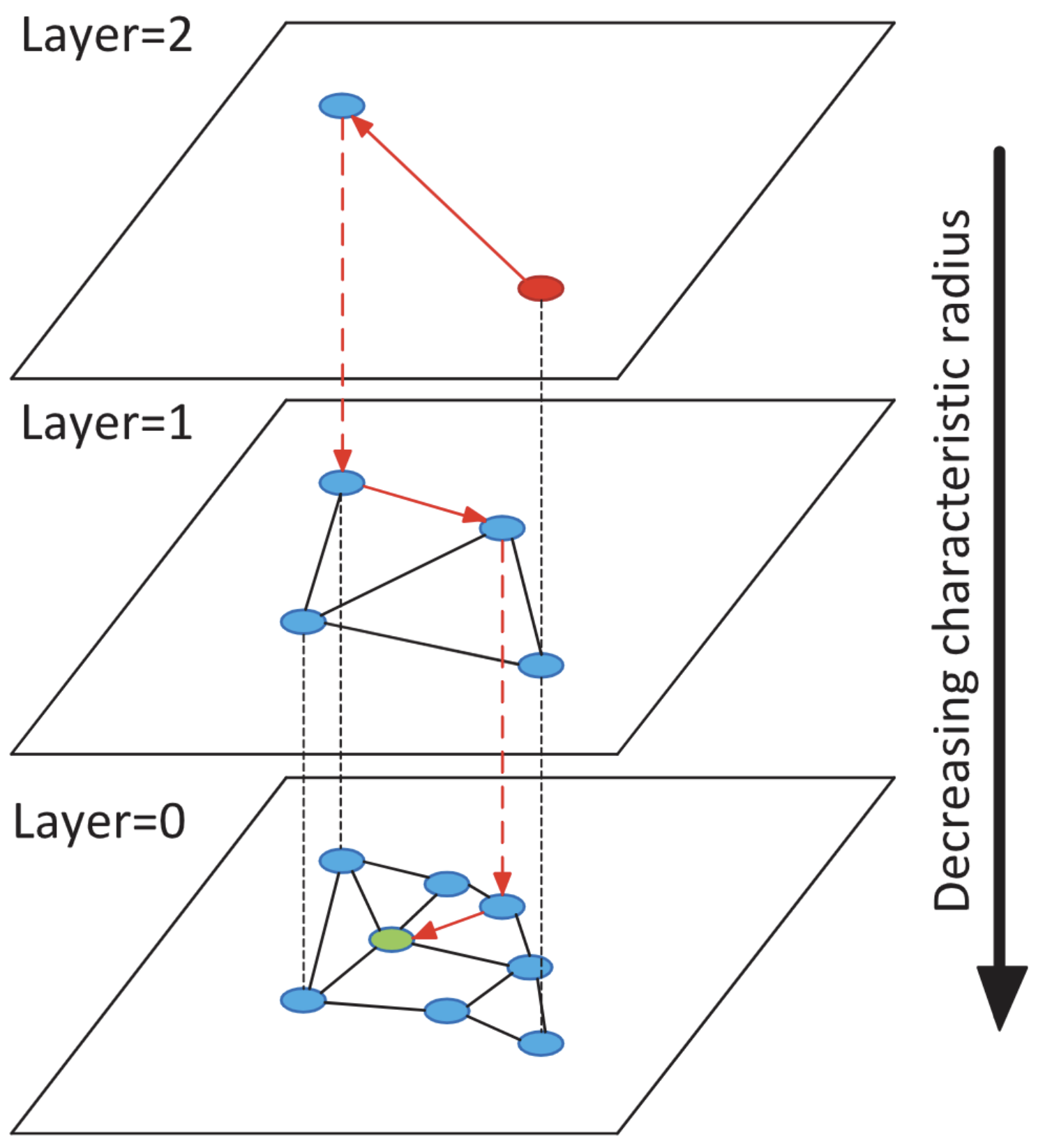
Vector Retrieval and GPU Acceleration
Exploration of the development of vector retrieval technology, transitioning from traditional CPU-based indexing methods to modern GPU-accelerated solutions. Highlights various classic algorithms in Approximate Nearest Neighbor Search (ANNS) and their applications in large-scale data processing, particularly focusing on their crucial role in fast semantic search and real-time decision support in the medical field.
Non-pathological image filtering in pathological multimodal data sets
Existing open-source pathology text-image paired datasets (e.g., Quilt-1m) are constructed by extracting frames from YouTube videos, though initial filtering strategies have been applied, significant noise (e.g., non-pathological images) remains. Training classifiers on datasets of varying scales and architectures demonstrates substantial performance disparities among different models. Experimental results further indicate that fine-tuning large models using an optimized dataset (filtered to exclude non-pathological data) significantly enhances their performance in downstream tasks.
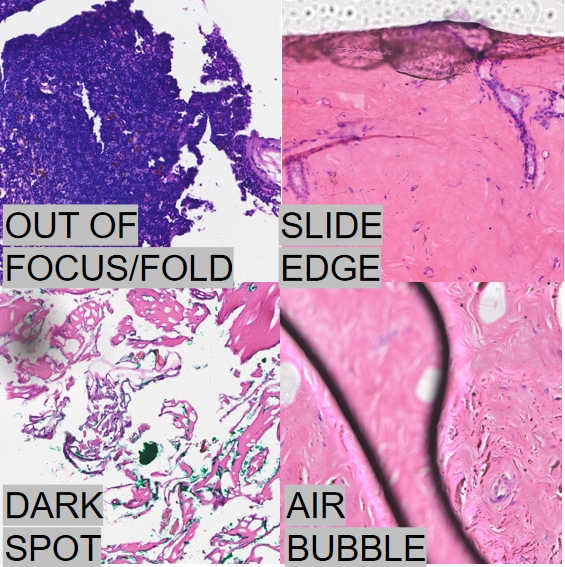
A Training Free Algorithm for Patch Level Quality Control in Pathological Images
The production process of pathological digital slides involves multiple critical steps, and potential quality issues in any of these steps may lead to defects such as image defocusing and tissue overlap. These abnormal regions result in the loss of pathological structural information, significantly compromising the accuracy and reliability of clinical diagnoses. Therefore, there is an urgent need to develop a rapid and efficient algorithmic framework to precisely identify and filter problematic regions, while further investigating the interference mechanisms and quantifying the impact of such low-quality image patches on the training of intelligent pathological analysis models.
Non-pathological Image Filtering
Data filtering can filter out data with high noise level
MLLM Evaluation in Breast Cancer
This project is focused on constructing a comprehensive benchmark to evaluate the performance of multimodal large language models (MLLMs) in breast cancer tasks.
Scaling law for pathology
The "Scaling Law for Pathology" project is aimed at exploring the efficacy of scaling laws within the specialized domain of pathology.
CLAM-based Image-Caption Generation
The high cost of labeling in the medical field has led to a lack of annotated data related to whole slide imaging (WSI), thereby limiting the performance of many downstream tasks, such as training and application of pathology CLIP.
KB-enhanced Pathology CLIP (public datasets)
KB-enhanced Pathology CLIP addresses the variability in performance of pathology foundation models across different branches of pathology.
Pathology Image-Caption Evaluation
Our research is focused on constructing a comprehensive benchmark to evaluate the performance
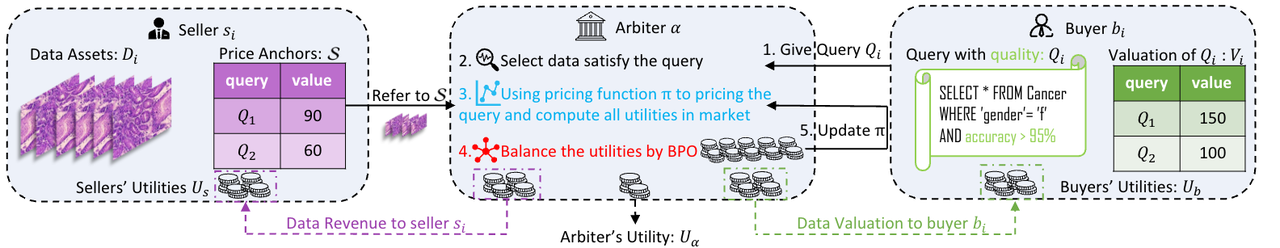
Image-Caption Data Market Demo
Acquiring high-quality training data is critical for the development of highly accurate and robust machine learning models, particularly as foundational models emerge.
GraphRAG Based Pathology LLM
Mining and analyzing private pathology data, and the rich information in the field of pathology is integrated by constructing refined pathology knowledge graph.
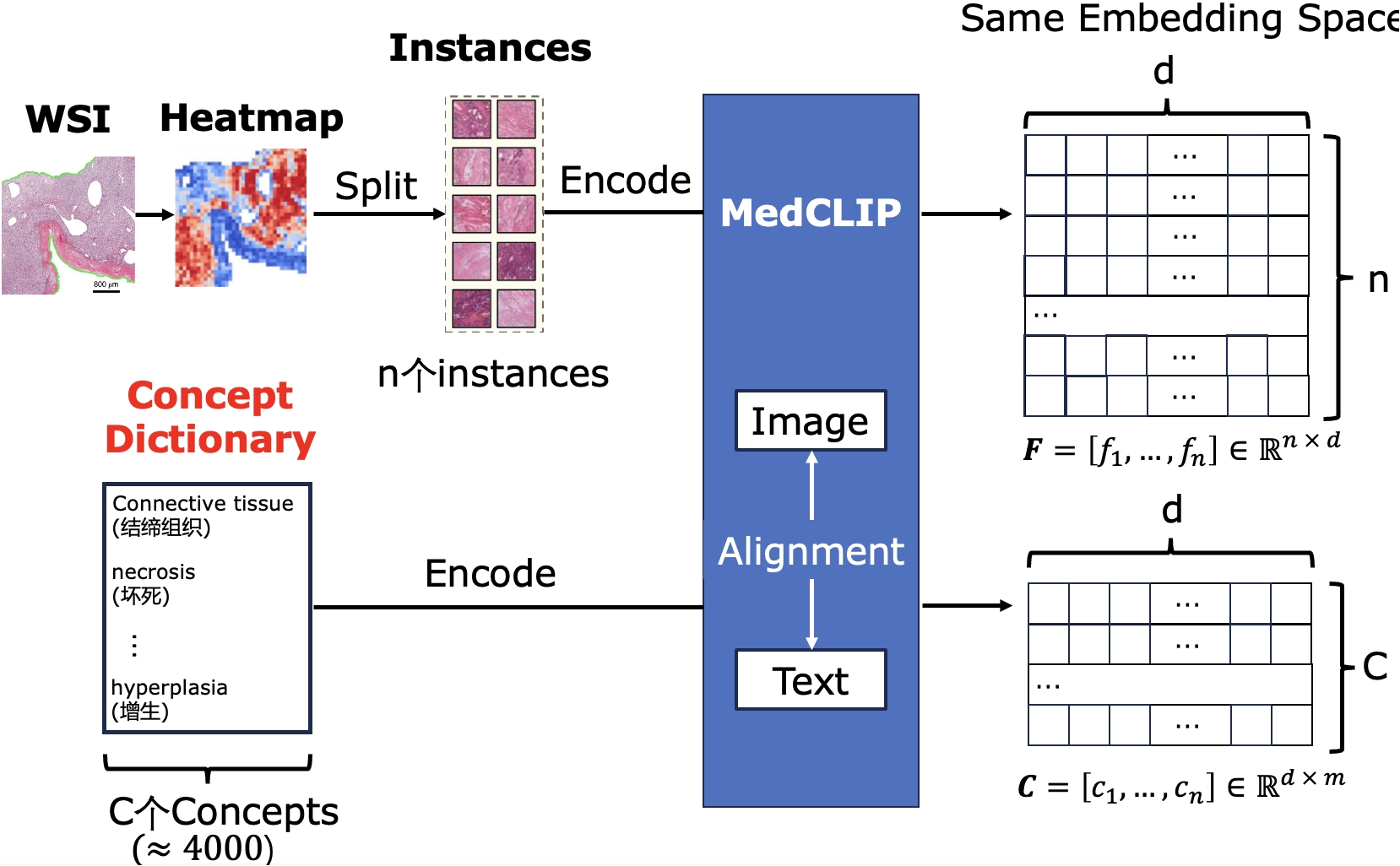
Pathology Image-Text Structured Alignment Based on Multiple Instance Learning
The pathology foundation model trained on massive pathological texts provides strong pathology image-text alignment capabilities.
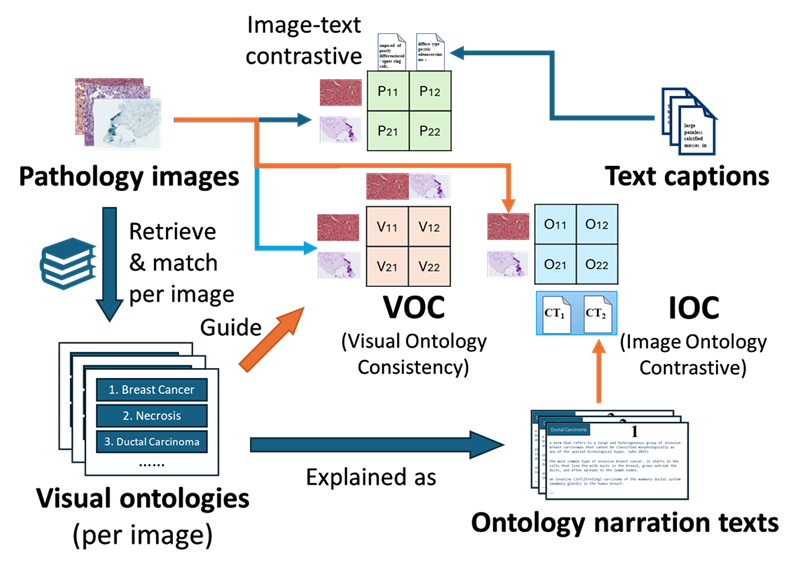
Pivot:Enhancing Pathology Image-Text Alignment with a Pathology Knowledge Base
Pivot aligns pathological images, pathological ontologies, and text.
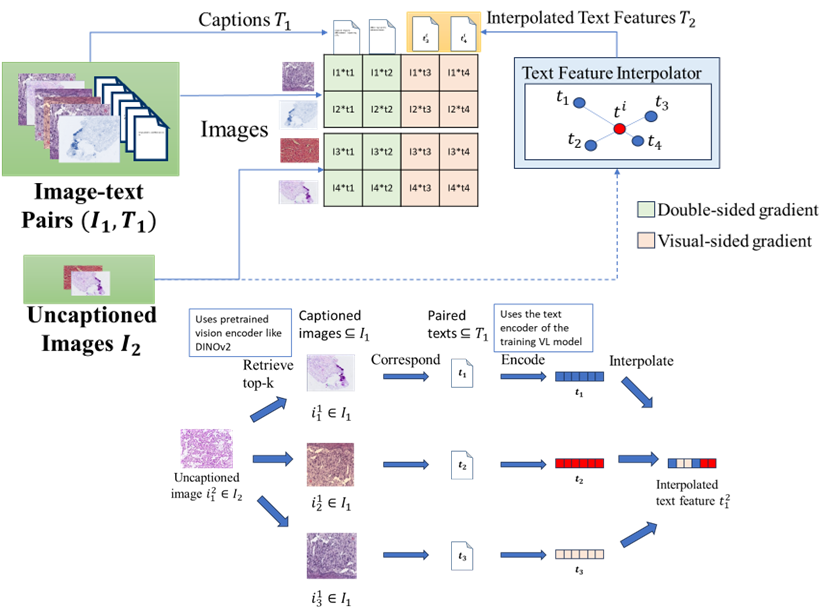
RIVL:Addressing Modality Missing in Pathology Image-Text Alignment Using Interpolation
Infering the semantic vector of the text annotation for a candidate image based on several images most similar to it.
Keywords
- Knowledge Graph
- Domain-specific
- Data Filtering
- Breast Cancer
- MLLM
- Visual-Language Model
- Labeling
- GraphRAG
- Data Market
- Balanced Pareto Optimization
- Multi-object Optimization
- Benchmark
- CLIP
- Scaling
- vector retrieval
- GPU acceleration
- ANNS
- Quality Control
- Training-Free
- Papers
- Show All


